Single-Cell APA Atlas of Abnormal Sperm
The SpermAPA Database is a specialized resource for exploring single-cell 3′ UTR alternative polyadenylation (APA) events in abnormal human sperm. It integrates high-resolution transcriptomic data to profile APA dynamics linked to spermatogenic defects and male infertility
Understanding APA in Sperm Abnormalities
Explore the molecular mechanisms linking alternative polyadenylation to sperm dysfunction
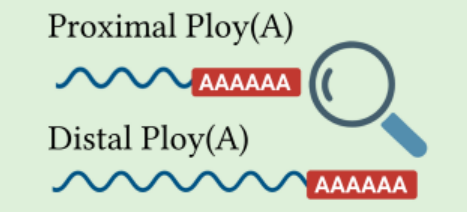
Alternative Polyadenylation
APA generates transcript isoforms with variable 3'UTR lengths, shaping mRNA stability and translation.
Dataset Integration
15 scRNA-seq datasets (3′ end library preparation, 10x Genomics) were obtained from GEO and GVA (GSE124263, GSE182786, GSE154535, GSE149512, HRA006558)
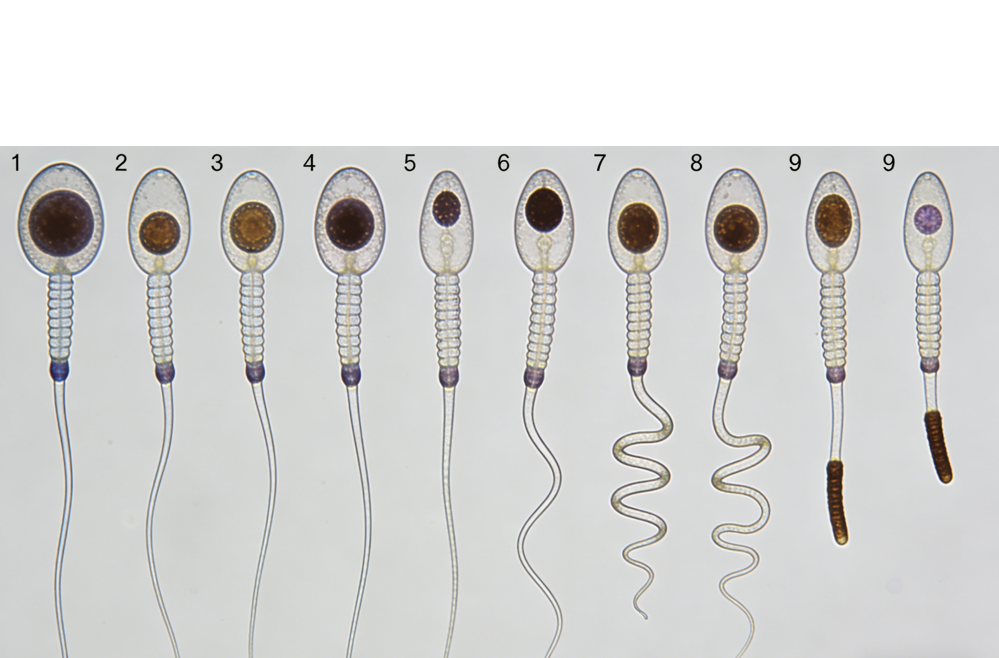
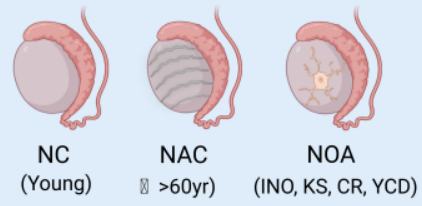
Sperm Abnormalities
Advanced paternal age compromises sperm quality and genetic integrity, whereas non-obstructive azoospermia (NOA) involves a complete or near-complete failure of spermatogenesis within the seminiferous tubules.

ML Classification
6 machine learning models (Logistic Regression, Random Forest, SVM, Naive Bayes, KNN, XGBoost) were used to predict azoospermia sample heterogeneity based on APA and gene expression (XGBoost AUC=0.97).